|
The Standard for the Exchange of Earthquake Data (SEED) is an international standard format for the exchange of digital seismological data. SEED was designed for use by the earthquake research community, primarily for the exchange between institutions of unprocessed earth motion data. It is a format for digital data measured at one point in space and at equal intervals of time. The SEED format consists of Volume Control Headers, Abbreviation Control Headers, Station Control Headers, Time Span Control Headers and finally Data Records. In complement to “Dataless” SEED volumes, exists the “Data-only” volume called Mini-SEED (see http://www.iris.edu for further information).
The purpose of these functions is to read and write miniSEED data files directly from Matlab, avoiding intermediate format conversion (like SAC or other formats for which many functions exist), having a full control on headers and formats. Note that IRIS proposes a complete C-library for miniSEED (libmseed) and there is a free Matlab toolbox using this library to read and write miniSEED files (see Stefan Mertl site), but it needs some compilation to be used (unsignificant for geeks, quite easy on UNIX/Linux, but no doubt more problematic for Windows users...). The present functions rdmseed.m and mkmseed.m are 100% Matlab code standalone scripts, slower, but fully portable and platform independent.
rdmseed: reading miniSEED file
Each data record is imported into a structure array, allowing to adress data blocks and header fields individually (useful for multi-channel files), just as concatenating all data with a simple cat(1,X.d) function. Time stamps are also converted into Matlab datenum format. The function reads miniSEED "data-only" using the two mostly used compression formats Steim-1 and Steim-2. General FDSN formats have also been implemented (ASCII, 16/24/32-bit integers, IEEE floats and doubles), and GEOSCOPE multiplexed old formats (24-bit, 16/3 or 16/4-bit gain ranged). All these formats should work but some of them have not been tested using real data. I also partly coded Steim-3 format but without a clear description and any file example... Since I never met any data file using this format, I don't know if it's really useful.
The function detects also automatically big/little-endian coded files.
Known Blockettes are 1000, 1001, 100, 500 and 2000. If there is no Blockette 1000 (which is mandatory in SEED format...), default 4096-byte record length, big-endian and Steim-1 compression are used. These values can be set using additional arguments.
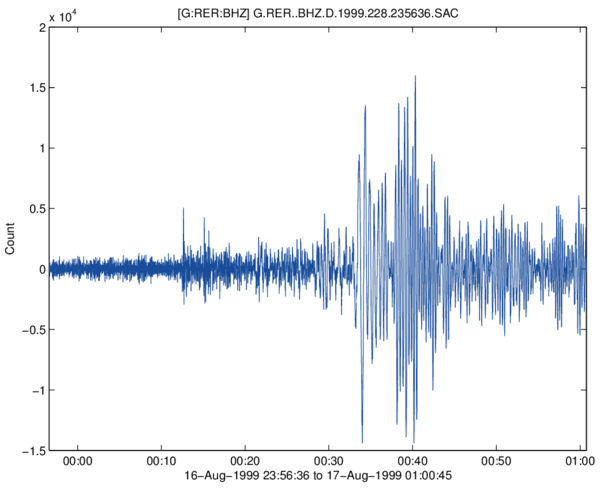
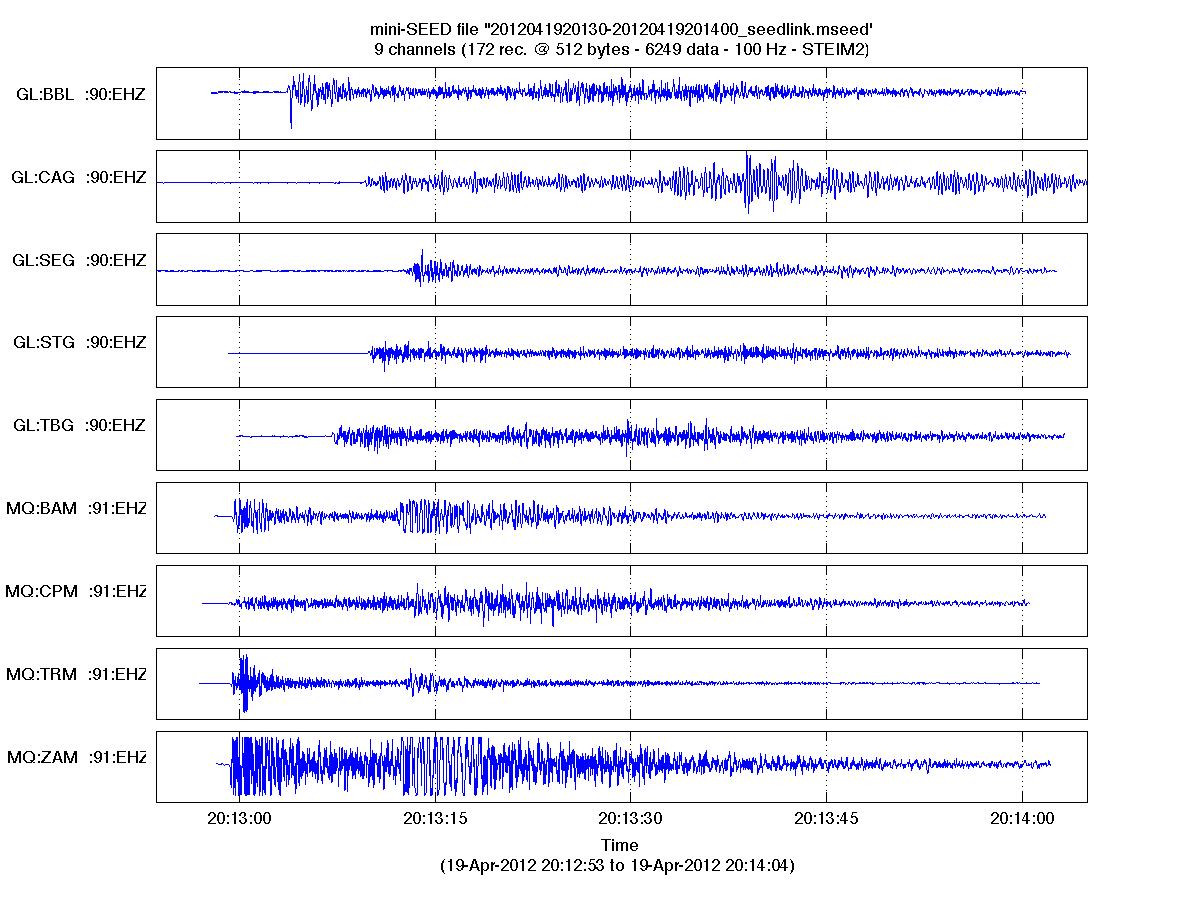
Using extra output argument, some analysis can be done on the data stream (detection of gaps and overlaps), and channel components are detected. Without any output arguments, or with an additionnal 'plot' input argument, the function plots the imported data in a new figure (works also in case of multi-channel file).
Steim-1/2 compression decoding strategy has been deeply optimized for Matlab. The proposed method, as vectorized as possible, is about 30 times faster than a 'C-like' loops coding... which is still 10 times slower than the same C-compiled program, but, well, this is the Matlab's other side of the coin!
mkmseed: writing miniSEED file
The function allows to export a data vector D to miniSEED file, giving origin date and time T0 and sampling rate FS (for monotonic data) or a time vector T. Header information is specified using the filename string with conventional naming "Network.Station.Location.Channel". Output file names will have appended ".Year.Day" and multiple file may be produced if data exceed a day.
Data encoding format can be specified (16/32-bit integers, IEEE float/double, Steim-1/2; Geoscope 16/3-4). If not, it will depend on the class of variable D. Note that D will be converted to relevant class depending on encoding format. Be aware that Steim-2 compression uses 30-bit signed integer to code difference between 2 consecutive samples; equivalent to a 29-bit coding. Steim-1 uses 32-bit signed integers to code differences, so a 31-bit equivalent. This means that full range of signed 32-bit integer data cannot be correctly encoded with these formats in some exceptional cases (may lead to data loss). Finally, Steim-1/2 encoding needs time processing to achieve a maximum compression ratio of about 6 in file size, while other formats are very fast but result in large files.
Binary file will be big-endian coded, with blockettes 1000, and default record length is 4096 bytes (this may be changed using input argument).
Type "doc rdmseed" or "doc mkmseed" for detailed usage.
RDMSEED Read miniSEED format file.
X = RDMSEED(F) reads file F and returns a M-by-1 structure X containing
M blocks ("data records") of a miniSEED file with headers, blockettes,
and data in dedicated fields, in particular, for each data block X(i):
t: time vector (DATENUM format)
d: data vector (double)
BLOCKETTES: existing blockettes (substructures)
Known blockettes are 100, 500, 1000, 1001 and 2000. Others will be
ignored with a warning message.
X = RDMSEED(F,ENCODINGFORMAT,WORDORDER,RECORDLENGTH), when file F does
not include the Blockette 1000 (like Seismic Handler outputs), specifies:
- ENCODINGFORMAT: FDSN code (see below); default is 10 = Steim-1;
- WORDORDER: 1 = big-endian (default), 0 = little-endian;
- RECORDLENGTH: must be a power of 2, at least 256 (default is 4096).
If the file contains Blockette 1000 (which is mandatory in the SEED
convention...), these 3 arguments are ignored.
X = RDMSEED without input argument opens user interface to select the
file from disk.
[X,I] = RDMSEED(...) returns a N-by-1 structure I with N the detected
number of different channels, and the following fields:
ChannelFullName: channel name,
XBlockIndex: channel's vector index into X,
ClockDrift: vector of time interval errors, in seconds,
between each data block (relative to sampling
period). This can be compared to "Max Clock Drift"
value of a Blockette 52.
= 0 in perfect case
< 0 tends to overlapping
> 0 tends to gapping
OverlapBlockIndex: index of blocks (into X) having a significant
overlap with previous block (less than 0.5
sampling period).
OverlapTime: time vector of overlapped blocks (DATENUM format).
GapBlockIndex: index of blocks (into X) having a significant gap
with next block (more than 0.5 sampling period).
GapTime: time vector of gapped blocks (DATENUM format).
RDMSEED(...) without output arguments plots the imported signal by
concatenating all the data records, in one single plot if single channel
is detected, or subplots for multi-channels file. Gaps are shown with
red stars, overlaps with green circles.
[...] = RDMSEED(F,...,'be') forces big-endian reading (overwrites the
automatic detection of endianness coding, which fails in some cases).
[...] = RDMSEED(F,...,'plot') forces the plot with output arguments.
[...] = RDMSEED(F,...,'v') uses verbose mode (displays additional
information and warnings when necessary). Use 'vv' for extras, 'vvv'
for debuging.
Some instructions for usage of the returned structure:
- to get concatenated time and data vectors from a single-channel file:
X = rdmseed(f,'plot');
t = cat(1,X.t);
d = cat(1,X.d);
- to get the list of channels in a multi-channel file:
[X,I] = rdmseed(f);
cat(1,I.ChannelFullName)
- to extract the station component i from a multi-channel file:
[X,I] = rdmseed(f);
k = I(i).XBlockIndex;
plot(cat(1,X(k).t),cat(1,X(k).d))
datetick('x')
title(I(i).ChannelFullName)
Known encoding formats are the following FDSN codes:
0: ASCII
1: 16-bit integer
2: 24-bit integer (untested)
3: 32-bit integer
4: IEEE float32
5: IEEE float64
10: Steim-1
11: Steim-2
12: GEOSCOPE 24-bit (untested)
13: GEOSCOPE 16/3-bit gain ranged
14: GEOSCOPE 16/4-bit gain ranged
19: Steim-3 (alpha and untested)
See also MKMSEED to export data in miniSEED format.
Author: François Beauducel <beauducel@ipgp.fr>
Institut de Physique du Globe de Paris
Created: 2010-09-17
Updated: 2014-05-07
Acknowledgments:
Ljupco Jordanovski, Jean-Marie Saurel, Mohamed Boubacar, Jonathan Berger,
Shahid Ullah, Wayne Crawford, Constanza Pardo.
References:
IRIS (2010), SEED Reference Manual: SEED Format Version 2.4, May 2010,
IFDSN/IRIS/USGS, http://www.iris.edu
Trabant C. (2010), libmseed: the Mini-SEED library, IRIS DMC.
Steim J.M. (1994), 'Steim' Compression, Quanterra Inc.
MKMSEED Write data in miniSEED file format.
MKMSEED(FILENAME,D,T0,FS) writes miniSEED file FILENAME from stritl
monotonic data vector D, time origin T0 (a scalar in Matlab datenum
compatible format) and sampling rate FS (in Hz). Encoding format will
depend on D variable class (see below).
MKMSEED(FILENAME,D,T,FS) where T is a time vector of the same length as
data vector D, will create data records of monotonic blocks of samples,
splitting each time the sampling frequency FS is not equal to time
difference between two successive samples (with a 50% tolerency).
Network, Station, Channel and Location codes will be extracted from FILENAME
which must respect the format "NN.SSSSS.LC.CCC" where:
NN = Network Code (2 characters max, see FDSN list)
SSSSS = Station identifier (5 char max)
LC = Location Code (2 char max)
CCC = Channel identifier (3 char max)
Final filename will have appended string ".YYYY.DDD" corresponding to year
and ordinal day of origin time (from T0 value). Multiple files may be
created if time span of data exceeds day limit.
MKMSEED(...,EF,RL) specifies encoding format EF and data record length RL
(in bytes). RL must be a power of 2 greater or equal to 256.
Data encoding format EF must be one of the following FDSN codes:
1 = 16-bit integer (default for class 2-bit, 8-bit, int16)
3 = 32-bit integer (default for class uint16, int32)
4 = IEEE float32 (default for class single)
5 = IEEE float64 (default for all other class)
10 = Steim-1 compression (D will be converted to int32)
11 = Steim-2 compression (D will be converted to int32)
13 = Geoscope 16-3 gain ranged (D will be converted to double)
14 = Geoscope 16-4 gain ranged (D will be converted to double)
MKMSEED(FILENAME,D) uses default value for T0 = now (present date and time),
and FS = 100 Hz.
File(s) will be coded big-endian, flags set to zero, blockette 1000, default
record length is 4096 bytes. Outputs have been tested with PQL II software
from IRIS PASSCAL (v2010.268).
See also RDMSEED function for miniSEED file reading.
Acknowledgments:
Florent Brenguier, Julien Vergoz, Constanza Pardo.
References:
IRIS (2010), SEED Reference Manual: SEED Format Version 2.4, May 2010,
IFDSN/IRIS/USGS, http://www.iris.edu
IRIS (2010), PQL II Quick Look trace viewing application, PASSCAL
Instrument Center, http://www.passcal.nmt.edu/
Author: François Beauducel <beauducel@ipgp.fr>
Institut de Physique du Globe de Paris
Created: 2011-10-19
Updated: 2014-05-07
: new version of rdmseed on April 21, 2012 with bug corrections (little-endian coding). Update your file!
Download rdmseed.m and mkmseed.m at Matlab Central file exchange (Copyright 2014 F. Beauducel / BSD License).
|